
|
1QKZ chain L |
|
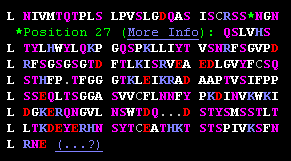
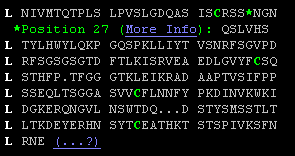
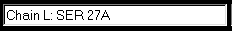
L NIVMTQTPLS LPVSLGDQAS ISCRSSQSLV L HSNGNTYLHW YLQKPGQSPK LLIYTVSNRF L SGVPDLRFSG SGSGTDFTLK ISRVEAEDLG L VYFCSQSTHF P-1-TFGGGTKL EIKRADAA L PTVSIFPPSSEQ LTSGGASVVC FLNNFYPK L DINVKWKIDGKE RQNGVLNSWT DQdskDST L YSMSSTLTLTKD EYERHNSYTC EATHKTST L SPIVKSFNRNE |
<Insertions at ...SSQSLVHSNGN... (27, 27A, 27B, ...) <Numbering gap: ...FP-TF... (P95 peptide bonded to T97) <Physical gap: residues without coordinates "dsk" |