
[Chapter 1:Chapter 2:Chapter 3:Chapter 4:Chapter 6][Top]
 | E-CELL2 User's Manual | Chapter 5: Creating User Defined Reactor Files [Chapter 1:Chapter 2:Chapter 3:Chapter 4:Chapter 6][Top] |
In the E-CELL2 System, cellular models can be created with the combination of three object types: Substance (to describe substance amount), Reactor (describe the reaction and the reactor system path), and System (to describe certain functions). Reactor describes the time-change of the Substance amount which plays an essential role under an E-CELL2 model.
Reactor objects can be freely manipulated by the user. Reactor objects must be compiled in separate with the E-CELL2 System and therefore must be converted to a format that E-CELL2 can read; if correctly written, the E-CELL2 System loads up the defined Reactor. For Reactor objects are written in C++, users can create almost any kind of a kinetic model.
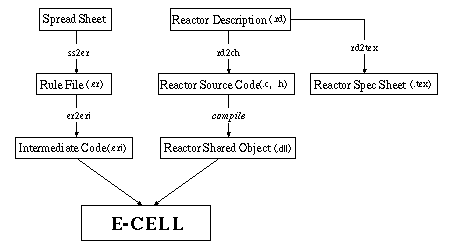
The graph above describes the files used in E-CELL2 System. As it can be seen, Reactors are created and compiled in separate of the E-CELL2 System itself. In order to create a Reactor object, the Reactor Description file (RD file) must be written with the actual commands and the required data(such as the equation and the parameters) inside. The RD file gets converted to C++ and .dll modules executable with E-CELL2 System. RD files can also be converted to a Reactor Spec Sheet in Latex format.
Two kinds of Reactors exist: Regular Reactors and Postern Reactors. While regular reactors use integrating functions to solve multi-derivative equations, postern reactors are separated from these integrating functions and used in cases where derivative equations are not suitable for calculating the Substance amount. The two reactors realize due to the multi-step calculation architecture; after the React step in the regular reactor is executed, the calculation will be performed in the Postern Step. Postern Reactors can be used in the following case:
While regular reactors calculate the reaction speed in order to define the substance amount, postern reactors do not consider the substance flux and handle all of the substance involved . Thus, while regular reactors use the velocity(Float v)method in performing calculations, postern Reactors use the SetQuantity(Float Q)method in returning calculation results.
Postern reactors have "PReactor" on the end of their name. (ex:
"ABCPReactor")
Each postern reactor must and can only handle a single substance where the substance cannot be defined more than once (if so, the simulation accuracy cannot be guaranteed). If multiple reactions effect a single substance, each reactions must be inside a singular postern reactor such as the GeneralRapid EquilibriumPReactor.
Though several General Reactors for simulation is already included in the E-CELL2 System Package (refer to section 5), in order to create non-standard reactions or to perform higher leveled simulations, it is required for the user to create their own Reactor files.
RD files can be created by including the keyword - value pair lines. Note for the following while creating RD files:
#" will be created
as comment lines. In order to use the # character at the start of the line
as a non-comment line, use \# or include a space before it. #s at non-starting
positions will be treated as normal characters. @CLASSNAME line is to define the Reactor Classname. The filename
of the Reactor MUST BE CLASSNAME.rd
in any case.
@BASECLASS line is to define the base class for that class. In
most cases, FluxReactor is defined.
@AUTHOR, @EMAIL, @DATE lines are to define those information into the reactor.
@BRIEF_DESCRIPTION line is a single-lined definition of the Reactor. The details
of these keywords are passed on to the E-CELL2 System.
@DESCRIPTION line is to define the full description of that Reactor.
@EQUATION line is to define the equation in LaTeX displaymath format.
Thus:
$$ Write Equation$$ \begin{displaymath} Write Equation \end{displaymath}
\[ Write Equation\] choose any of the above.
%SUBSTANCE line is to define the Substance from Substrate, Product,
Catalyst, and Effector, its maximum amount, minimum amount and their comments
separated by commas ",". The max/min amounts can be defined in decimal
integers, but it is also possible to set the maximum as "Inf" for
infinite. Comments can be used as a section to describe special Substances.
As of now, the information here will only be reflected to the Spec Sheet.
Ex: "%SUBSTANCE:Substrate, 1, 10" means that the amount of
the substrate is between 1 and 10.
@NOTES line is to define any comments that need to be noted.
%PARAMETER line is to define the parameters that become the argument
for that Reactor in parameter name, type, unit, comment format separated by
commas ",". Input "Int" or "Float" as parameter
type according to the value (which are NOT identical to the "int"s
and "float"s used for C++). "Int"s in E-CELL2 Windows version
are 32bit integers and "Float"s are 80bit firm decimals (but because
of JNI(Java Native Interface), the accuracy is 64 bit in the GUI version and
80bit in the batch version)
%INCLUDE_FILE_H line is to define the .h filename to be included
in the reactor. Do not embrace file names with < > or " ".
@PRIVATE line is to define private items used.
@PROTECTED line is to define restricted items used.
@PUBLIC line is to define the public items used.
If FluxReactor.h is included in the file, Reactor.h, Reactant.h, and RootSystem.h are all automatically included in the file are need not to be included. Note that Stepper.h used in E-CELL1 is not used in E-CELL2. If not including FluxReactor.h, including StandardHeaders.h is desired in ordered to include the former three header files.
@OPTION_C line is to define new methods and macros. The data here will
be appended to the top of the .cpp file.
@INITIALIZE_FUNC line is to define the C++ code to be executed
only once at the initialization point. This line can be used to set the non-changing
elements and to check parameter ranges.
@REACT_FUNC line is to define the process that must be done for each
calculation step. As default:
if including the FluxReactor, process() method can be used to easily
perform the 2. task. Argument for the process() method is the molecular
reacting amount per second(in Float).
The activity that the reactor handle is the value per single step, but the value shown in the reactor window are values per second.
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| (*1) will be treated as comments (*2)Dependent on the Reactor |
Figure: Method list 2.2 Reactor Description ExampleIn order to create a Reactor file for the kinetic equation such as one below:
we shall name the Reaction "MichaelisUniUniReversibleReactor" and the keyword definition will be such as the following:
In the @REACT_FUNC line, variable S must be declared to describe Substrate concentration. Thus: P and E must identically be declared. 3 Conversion to Reactor Source Code (.C .h file) and DLLTo create Reactor source and executable (DLL) modules for E-CELL2 system, first press the Reactor tab of the ModelingLauncher. By selecting the .rd for input and pressing the "Execute" button, the launcher will automatically create the source code and DLL files by compiling. Refer to the ModelingLauncher tutorial in Chapter 6 for more details.  Figure: Modeling Launcher 4 Load the E-CELL2 SystemIn order for E-CELL2 to be executed, the following precautions must be noted. The Reactor (dll files) defined in the Rule must be placed under the Reactor Directory (As default, DLLR and DLLRB are set in ECELL2.BAT and ECELL2BB.BAT). Normally the created reactor dlls will be placed in these directories. E-CELL2 will terminate with an error message if the designated Reactor itself or the path do not exist. 5 General Reactors from the E-CELL2 Project
Figure: Standard Reactor
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||